On Habrahabr in the beginning of 2013 after the announcement of the launch of the European mega-project to study the human brain with a budget of more than a billion euros, a 10-year, was published by the corresponding
Notes . At the end of last year, the same project was officially launched, and marked the first means, but still was not written a single word of what scientific basis is the basis for the forthcoming titanic work, comparable in importance and scale with decoding the human genome and a manned mission to Mars.
At the end of the post you can just ask questions of the person directly involved in the team The Blue Brain Project, the answers to which will leave a separate post. B> i>
Instead of preface h4>
So, all the mega-project is divided into 12 sub-projects, which are, like, and sold separately - as here cross the high-load computing and biology, for example - but closely related and intertwined. I will not bore long and boring stories about each of the projects. To do this, there are short videos on the official Youtube-канале The Human Brain Project (HBP, additional information can be found at official site ). The overall goals and objectives of the project are indicated in this video:
In my opinion, one of the basic, fundamental and therefore the most important subproject is the direct study of the structure of neurons, their contacts and location - neural network - a brain (Subproject 1 - The Mouse Brain Subproject), as well as the visualization of such networks for the subsequent construction of models and actually implement this knowledge in the gland, in the calculations.
Let's start with the minute stories. The exact date of the opening of the nerve cells, neurons , as such, perhaps, can not lead. We can only say with certainty that by the end of the 19th century had accumulated sufficient knowledge about the functioning of the nervous tissue to gentlemen in 1906 Golgi and Рамон-и-Кахаль shared the Nobel Prize in Medicine for his work on the structure of the nervous system and the classification of nerve cells.
Over the last hundred years we have understood the basic principles of the functioning of the nervous tissue, learned even the relatively primitive level to intervene in the processes occurring in it (eg, anesthesia or surgery affecting directly the nervous tissue), but until now, a large spot had it exactly how the brain stores information as it specifically neurons processed, converted.
How do I know where they are going and where the signals in the nervous tissue? H4>
Omitting the details of that nerve cells are different, they are in many ways linked to the nervous tissue, have different functions, it is possible, however, to identify common features in the structure of neurons:
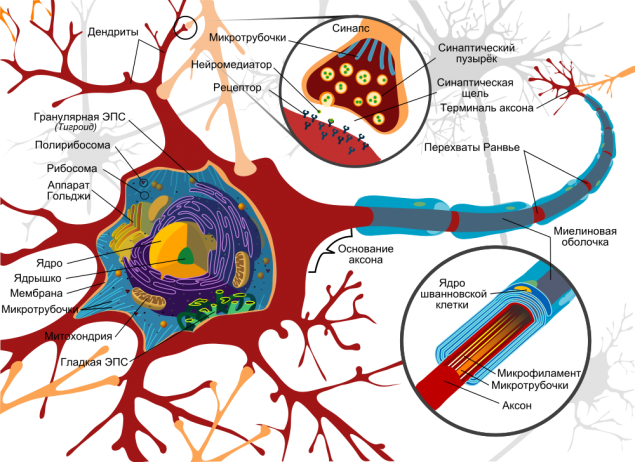
The model neuron. Источник
Serve basis signaling axons as electric cables which extend from one neuron to another and required for the transmission of pulses. Theoretically, they can achieve great length - up to one meter. Are attached to the same axons of other neurons via synapses, while at the end of each axon synaptic vesicles are neurotransmitters:
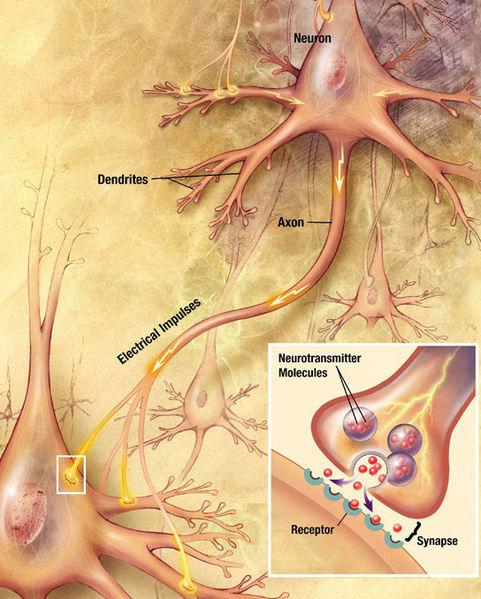
momentum transfer between two neurons. Source i>
Thus, the first thing we are interested in two things: the axon itself and attaching the axon to the next nerve cell, which can be determined by synaptic vesicles with neurotransmitters. For this, we are only two methods, by and large: fluorescent optical microscopy and electron microscopy.
Anticipating a logical question, why I do not consider MRI (магнитно-резонансную tomography ) and functional MRI, these methods are used more in order to localize the areas responsible for certain functions (hearing, sight, etc.), but do not see the individual neurons and their networks.
Next, I'll talk about the first subproject - the study of mouse brain, but not human (unfortunately, ethical reasons prevail over reason). The second sub-project dedicated to just use MRI. Thus, the study of the brain of the poor mice give us information about the fine structure of the brain, which then scientists have been trying to contact by MRI.
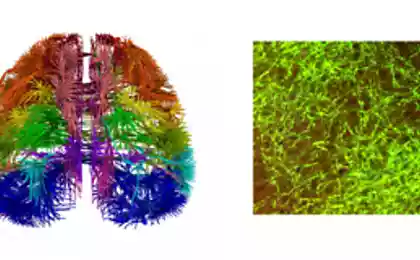
But back. In the first case - the case of fluorescence microscopy - need to paint the neurons with a fluorescent dye, and then get a picture of the sample at the excitation of the dye with light at a specific wavelength (UV often), but resolution of this method will only see themselves axons, for example, or only dendrites, so they are painted in different colors, but overall we had a hard can determine how neurons interact.
A few examples:
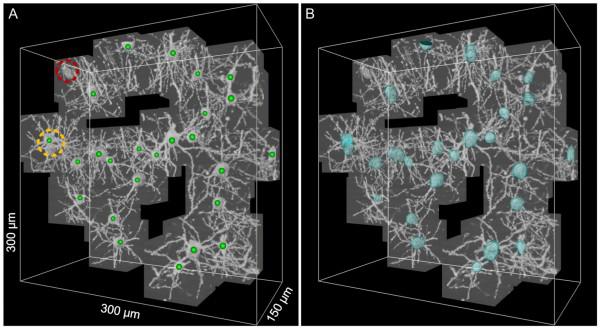
A new milestone - Fluorescence 3D-microscopy. More on Russian i>
Video English how fluorescent microscopy works in the case of nerve cells can be found тут. Unfortunately, there is no embedded player, so go to the link, please ...
Another method - is electron microscopy, the resolution of which is a few nanometers in the case of a nanometer scanning and in case of transmission. However, these methods are suitable for any surface analysis (scanning) or thin specimens - to 100 nm (translucent). It seems that we hit a dead end, is not it ?!
The catch is compounded by the fact that the entire tissue (brain or any other) for the most part consists of three main elements: carbon, nitrogen and oxygen, that is, the light elements. The contrast between the light elements - and even blended and evenly distributed - not give even the most advanced in the world of the electron microscope. It's not physically possible.
Long or short it ... but scientists have found a way out of this trap. What information do we want to extract? In fact, we need to know the location of the cell membranes within the tissue, and then we could "restore" the location of the cells themselves and their "internal". And the solution was found in the use of salts of heavy metals to "touch-up" membranes. For example, it was found that the compound of osmium - OsO 4 sub> - very well deposited on the cell membranes or as it is commonly called, is concentrated.
Actually, the case for small - to visualize. For a long time only used transmission electron microscopy, which requires very thin samples that prompted engineers to develop ultra-cryo-microtome that can be cut from the fabric layer to 30-50 nm. Transmission electron microscopy allowed many generations of scientists to study the brain slices, gave us knowledge about the internal structure of the cells, was used to study the bio-tolerance implants, but today this method is replaced by another, scanning electron microscopy with 3D reconstruction.
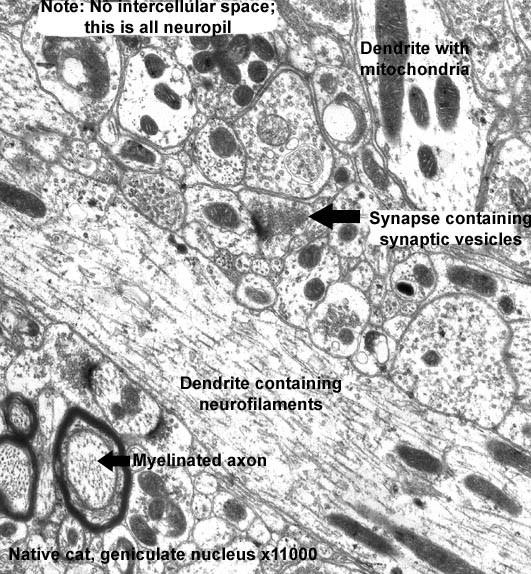
TEM-micrograph neuropil - accumulation processes of nerve cells (an increase of 11 000-fold). Источник
3D microscopy: new opportunities h4>
Okay. One way or another we have been able in the chaos of nerve cells (and it really is chaos, because in past neurons themselves in a variety of supporting tissue cells) to identify the point of articulation of individual neurons, but in terms of building neural networks, it is actually useless information, because there is no the opportunity to see the location of cells in three-dimensional space, that is, information about the three-dimensional organization is hidden from us. And there was found, I would say, a unique method - three-dimensional microscopy, which has become possible only in the past few years, thanks to the development of computing power and processing electronics microscopes.
In center electron microscopy EPFL has been successfully applied approach combines the full cycle of brain tissue sample preparation and analysis, followed by its 3D reconstruction. Данное video presents the main stages of the experiment:
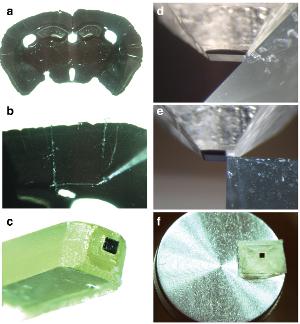

Results of fixing the water sample and a special resin substitution (1:30 video): a. Mouse brain slice thickness of 80 microns, bc. Cut-out and fixation area of interest 3x3 mm, df. The final treatment with a microtome. I>
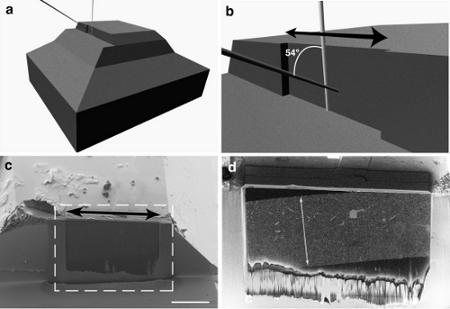

The principle of scanning electron microscopy and focused ion beam FIB / SEM (4:00 video): ab. A schematic arrangement of FIB-beam shear of the fabric and the electron beam, giving the image, cd. SEM-micrographs of the treated tissue.
I>
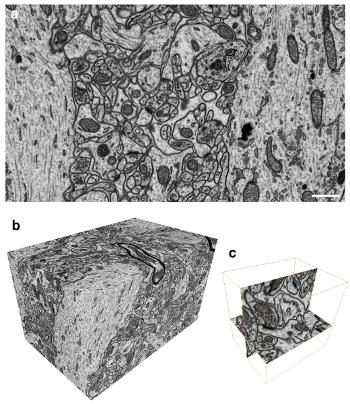
a. Imaging with FIB / SEM (5:55 in the video) and bc. The end result (8:00 in the video) i>
The main advantage of the method - "cutting" only 5 nm brain tissue layer by a focused ion beam (FIB) and then scanning for imaging an electron beam (SEM). Thus, we are able to very accurately examine the structure of the nervous tissue, but more importantly - now the processing of such stacks (up to 2000 images) can be performed virtually automatically, without human intervention, through the use of special algorithms for image analysis, offering accurate reconstruction often no less than professionals, microscopists.
Group Mark Canton ( Marco Cantoni ) used a free-ware program specifically designed for this kind of analysis - ilastik . The project has its own representation at github , so if someone from dear readers Habra will be interesting to participate in the project or to offer their ideas, it is not I think that the developers will be denied.
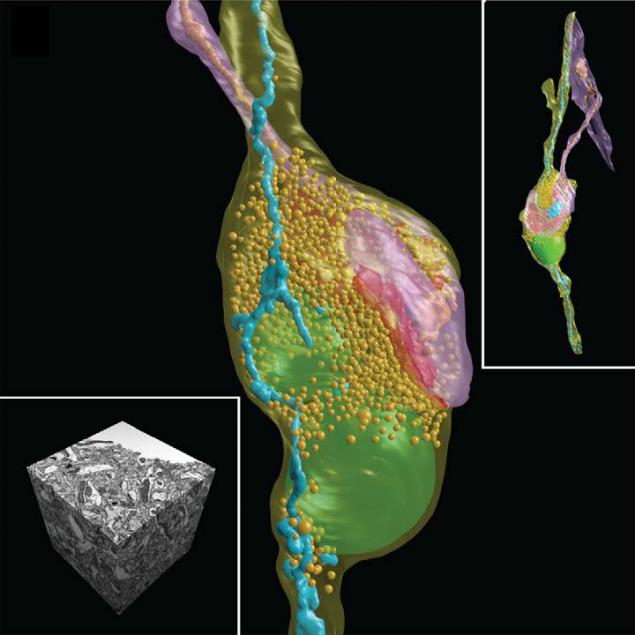
In the end, we have a 3D map of a piece of brain in the form of a cube with an edge of only a few tens of microns (yes, a little, but Moscow is also not built in a day), but do not despair and give up. We got almost what they wanted - quite precisely visualized in 3D is not the whole neural network, of course, but the individual neurons, and one shazhochek closer to the main goal. This is according to the authors of an ambitious project to better understand the principles of organization and functioning of the brain.
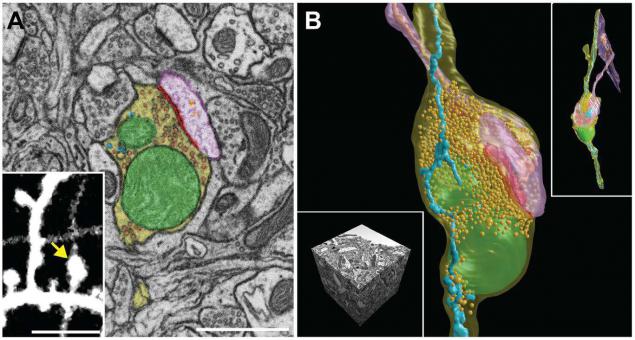
SEM-micrographs and 3D reconstruction of synoptic contact and all the membranes inside the neurons in the brain of the mouse. Scale A - 1 micron, insert the image A - 5 microns. I>
PS: Looking back and realizing what a jerk electron microscopy made over the last decade, I would like to point out that automated systems, including those developed under this project will allow in 5-7 years to handle much larger data sets, and then the case will remain for small - only to create a 3D map of the brain or "device, through which we think, what we think" (Ambrose Bierce).
When training materials were used open source: 1, 2 , 3
Source: habrahabr.ru/post/211997/